This represented a major change in the Cytoscape architecture it is a more modularized, expandable and maintainable version of the software. The Cytoscape core developer team continues to work on this project and released Cytoscape 3.0 in 2013. Version 3.0 was released Feb 1, 2013, and the latest version, 3.4.0, was released in May 2016. Version 2.0 was initially released in 2004 Cytoscape 2.83, the final 2.xx version, was released in May 2012. Default annotations and ontologies date from August 2010 and will be updated irregularly. Version 1.1.1 is the last stable release for the 1.0 series. Current version : BiNGO 3.0.3 (compatible with Cytoscape 3.0 and above) Notice: using default annotations and ontologies is no longer recommended for anything more than a quick-and-dirty analysis. Cytoscape was initially made public in July, 2002 (v0.8) the second release (v0.9) was in November, 2002, and v1.0 was released in March 2003. Now, it is developed by an international consortium of open source developers. Install Necessary Plugins Start Cytoscape Under the Plugins menu, select Manage Plugins In the Search dialog, type PSICQUIC Find PSICQUICUniversalClient plugin and if it is not already installed, click the Install button. Cytoscape also has a JavaScript-centric sister project named Cytoscape.js that can be used to analyse and visualise graphs in JavaScript environments, like a browser.Ĭytoscape was originally created at the Institute of Systems Biology in Seattle in 2002. In this tutorial, you will learn how to import interactions from public repositories. Plugins may be developed using the Cytoscape open Java software architecture by anyone and plugin community development is encouraged. Plugins are available for network and molecular profiling analyses, new layouts, additional file format support and connection with databases and searching in large networks. Such functional categories are typically derived. Given a list of genes resulting from an experiment, enrichment analysis enables to identify functional categories that are over-represented. Additional features are available as plugins. Enrichment analysis (also known as functional enrichment) is an helpful technique for high-throughput data interpretation. 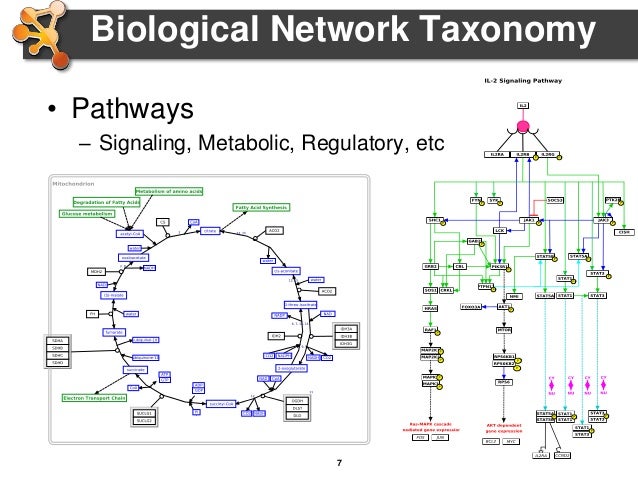
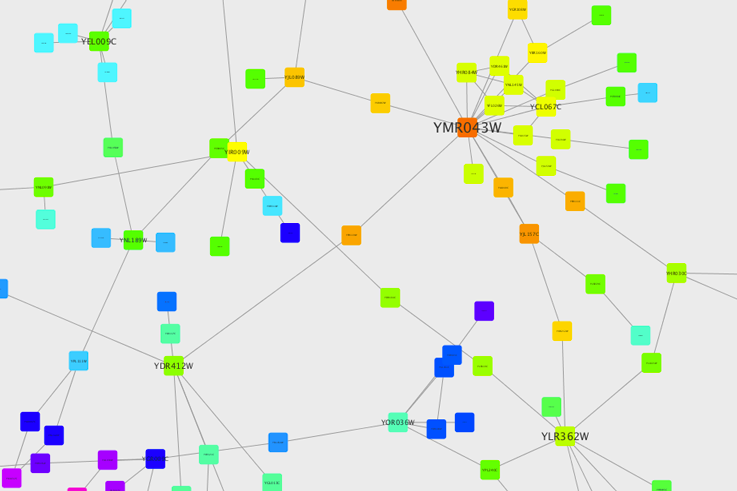
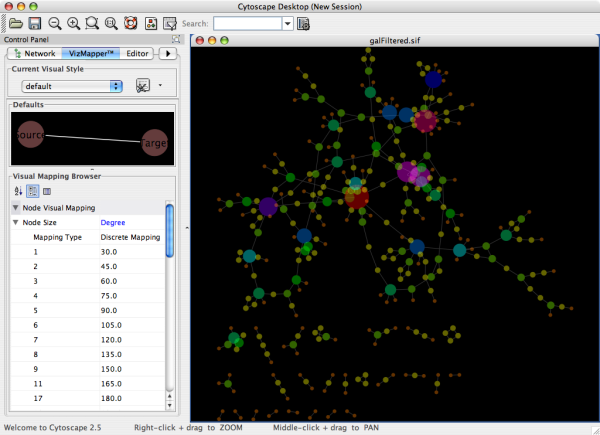
Cytoscape is an open source bioinformatics software platform for visualizing molecular interaction networks and integrating with gene expression profiles and other state data. add_edge ( sour, targ, key = key ) graph. 
copy () sour = d targ = d if multigraph : key = d. update ( node_data ) for d in data : edge_data = d. get ( "data" )) for d in data : node_data = d. get ( "directed" ) if multigraph : graph = nx. NetworkXError ( "name and ident must be different." ) multigraph = data. To install the KPM plug-in with Cytoscape 3 and above, go to the Apps -> App manager and seach for. Cytoscape user's manual: Examples - > G = nx.path_graph(2) > nx.cytoscape_data(G) # doctest: +SKIP )]) """ if name = ident : raise nx. In order to include edges, there are several options which can be found on the Cytoscape.js page under traversing such as nodes.edgesWith (), nodes.edgesTo (), nnectedEdges (), etc. Installation Download and install Cytoscape. See Also - cytoscape_graph: convert a dictionary in cyjs format to a graph References. Raises - NetworkXError If the values for `name` and `ident` are identical. Returns - data: dict A dictionary with cyjs formatted data. ident : string A string which is mapped to the 'id' node element in cyjs format. Parameters - G : NetworkX Graph The graph to convert to cytoscape format name : string A string which is mapped to the 'name' node element in cyjs format. Def cytoscape_data ( G, name = "name", ident = "id" ): """Returns data in Cytoscape JSON format (cyjs). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
